JCRB1095 KYSE150
Cell information
Important Notice(s)Announcement of unavailability of KYSE series cell lines to oversea users outside of Japan
Cell type:general cells (View Pricing Information)
| JCRB No. | JCRB1095 | Cell Name | KYSE150 |
|---|---|---|---|
| Profile | Poorly differentiated squamous carcinoma cell line from human esophageal cancer. | Other Name | |
| Animal | human | Strain | |
| Genus | Homo | Species | sapiens |
| Sex | F | Age | 49 |
| Identity | available | Tissue for Primary Cancer | esophagus |
| Case history | esophageal cancer | Metastasis | |
| Tissue Metastasized | Genetics | modal chromosome number = 84XX | |
| Life Span | infinite | Crisis PDL | |
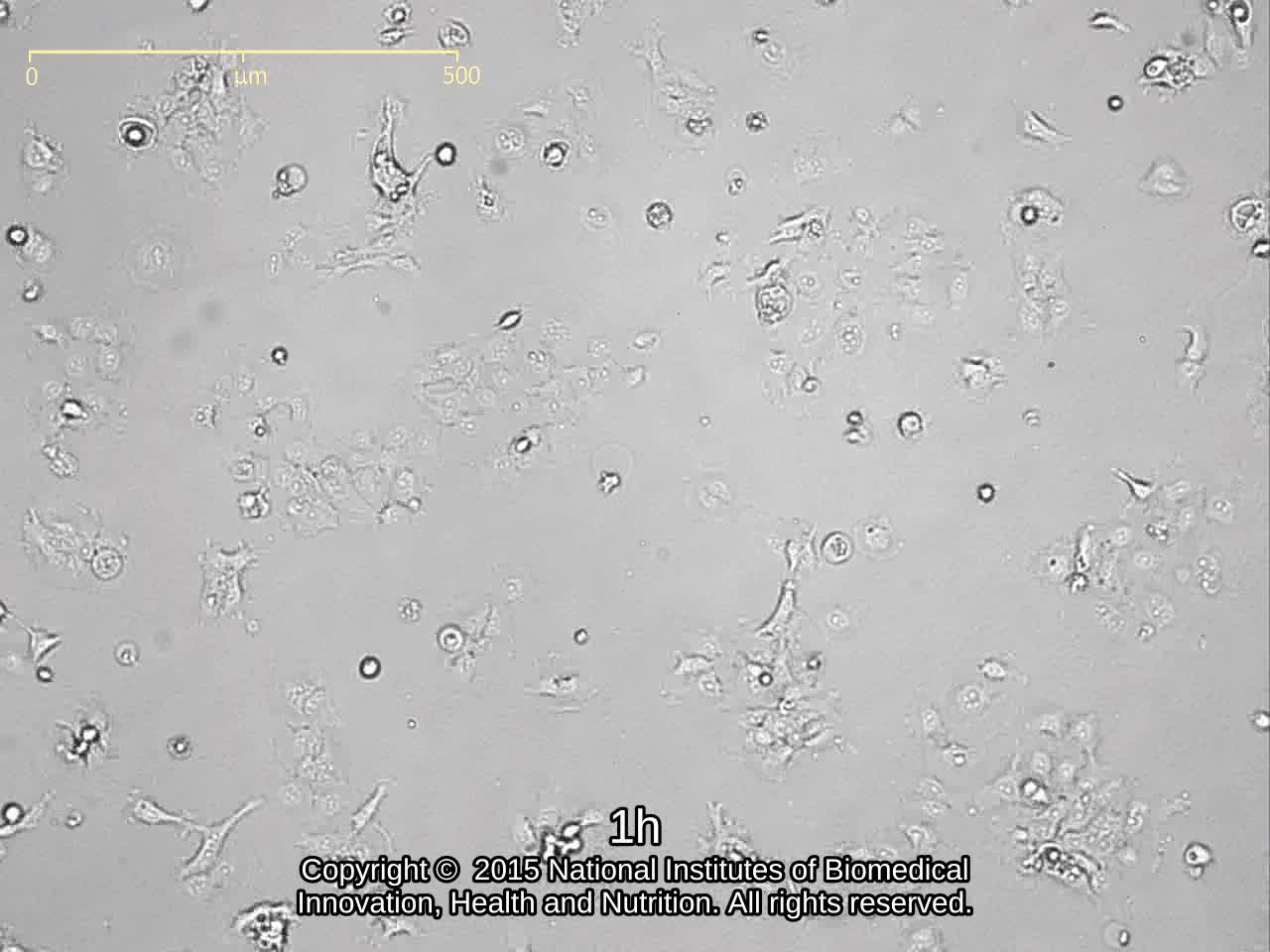
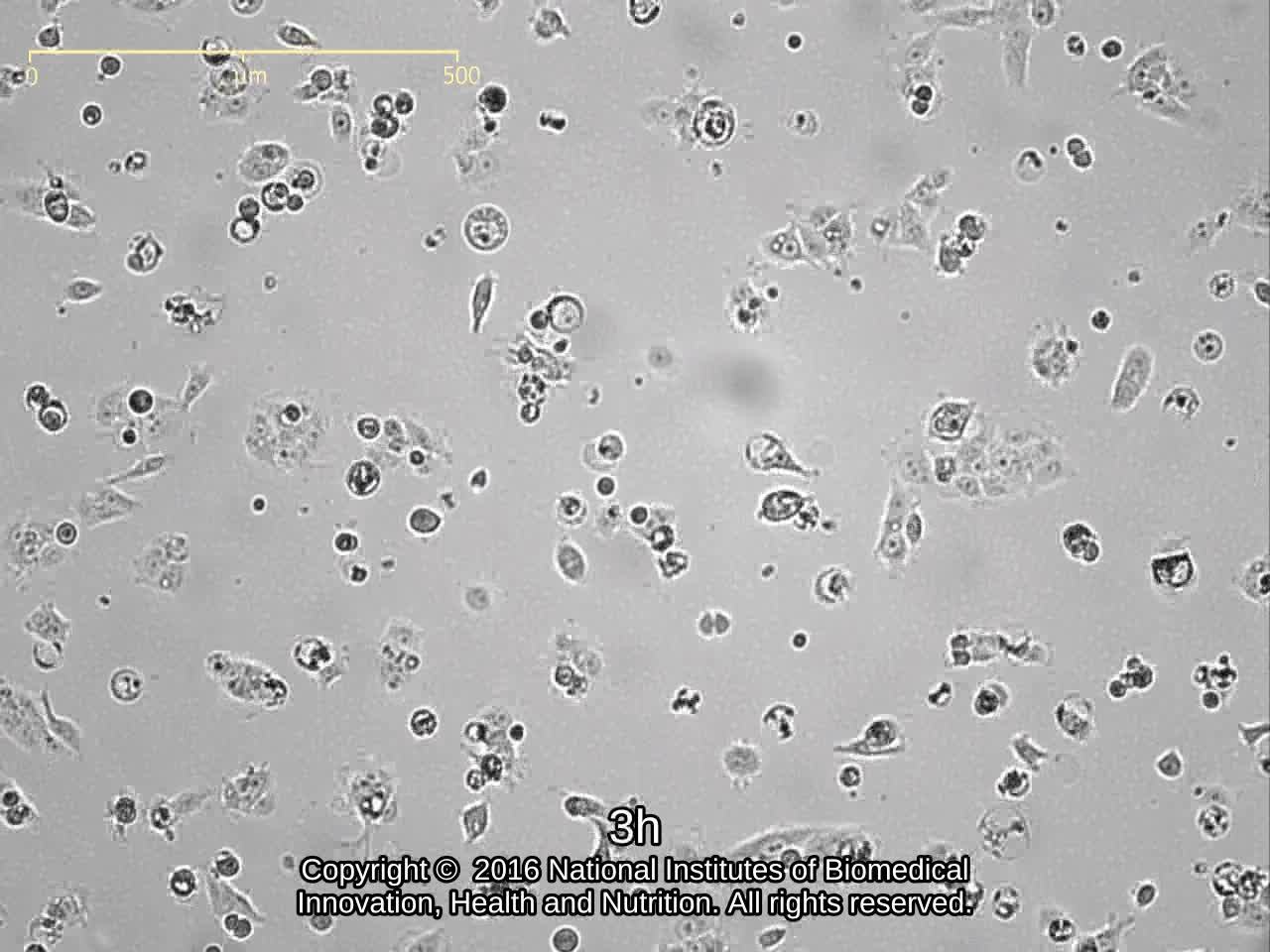
| Morphology | epithelial-like | Character | |
| Classify | tumor | Established by | Shimada,Y. et al. |
| Registered by | Shimada,Y. | Regulation for Distribution | |
| Comment | Year | 2003 | |
| Medium | Ham's F12 medium with 2% fetal bovine serum. | Methods for Passages | |
| Cell Number on Passage | Race | Japanese | |
| CO2 Conc. | 5 % | Tissue Sampling | esophagus |
| Tissue Type | poorly differentiated squamous carcinoma |
| Detection of virus genome fragment by Real-time PCR | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Detected DNA Virus | tested | Detected RNA Virus | tested | ||||||
| CMV | - | parvoB19 | - | HCV | - | HTLV-1 | - | ||
| EBV | - | HBV | - | HIV-1 | - | HTLV-2 | - | ||
| HHV6 | - | HTLV-1 | - | HIV-2 | - | HAV | - | ||
| HHV7 | - | HTLV-2 | - |
-/negative. +/positive. nt/not tested. (positive (+) does not immediately mean the production of infectious viral particles.) |
|||||
| BKV | - | HIV-1 | - | ||||||
| JCV | - | HIV-2 | - | ||||||
| ADV | - | HPV18 | - | ||||||
| Notes | |||||||||
| Reference | |
|---|---|
| Pubmed id:11520067 | Gene expression profiling in human esophageal cancers using cDNA microarray. Kan T,Shimada Y,Sato F,Maeda M,Kawabe A,Kaganoi J,Itami A,Yamasaki S,Imamura M Biochem Biophys Res Commun. 2001 Aug 31;286(4):792-801 |
| Pubmed id:10995879 | PTEN/MMAC1 expression in esophageal squamous cell carcinomas. Ding Y,Shimada Y,Kano M,Itami A,Kawabe A,Maeda M,Li Z,Hong T,Sato F,Kaganoi J,Imamura M Int J Oncol. 2000 Oct;17(4):695-9 |
| Pubmed id:10362086 | Induction of human leukocyte antigen (HLA)-A2-restricted and MAGE-3-gene-derived peptide-specific cytolytic T lymphocytes using cultured dendritic cells from an HLA-A2 esophageal cancer patient. Kanaoka S,Yamasaki S,Okino T,Inoue N,Shimada Y,Kaneko M,Otaka A,Fujii N,Imamura M J Surg Oncol. 1999 May;71(1):16-21 |
| Pubmed id:8486532 | Hyperthermic enhancement of cytotoxicity and increased uptake of cis-diamminedichloroplatinum(II) in cultured human esophageal cancer cells. Miyahara T,Ueda K,Akaboshi M,Shimada Y,Imamura M,Utsumi H Jpn J Cancer Res. 1993 Mar;84(3):336-40 |
| Images |
|---|
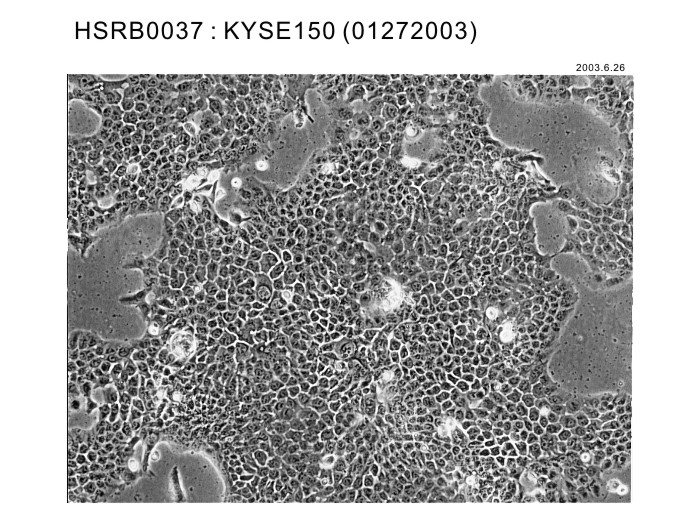   |
| Movies |
|---|


|
LOT Information
Viability/Growth rate/Cell number are represented as actual values measured at lot presentation in JCRB, but are not guaranteed values. Additionally, the doubling time is a rough value measured during passages.| Cell No. | JCRB1095 | Cell Name | KYSE150 |
|---|---|---|---|
| LOT No. | 12112008 | Lot Specification | distribution |
| Medium | Ham's F12 medium with 2% fetal bovine serum | Temperature | 37 C |
| Cell Density at Seeding | 6 - 8 x 10^4 cells/ml | Methods for Passages | 0.25% trypsin and 0.02% EDTA |
| Doubling Time | approx. 2 days | Cell Number in Vial (cells/1ml) | 1.1 x 10^6 |
| Viability at cell freezing (%) | 95 | Antibiotics Used | free (pretreated with MC-210 and BM cycline) |
| Passage Number | Unknown (10 at bank) | PDL | |
| Sterility: MYCOPLASMA | - | Sterility: BACTERIA | - |
| Sterility: FUNGI | - | Isozyme Analysis | Confirmed as human by NP, G6PD (type B), MD. |
| Chromosome Mode | Chromosome Information | ||
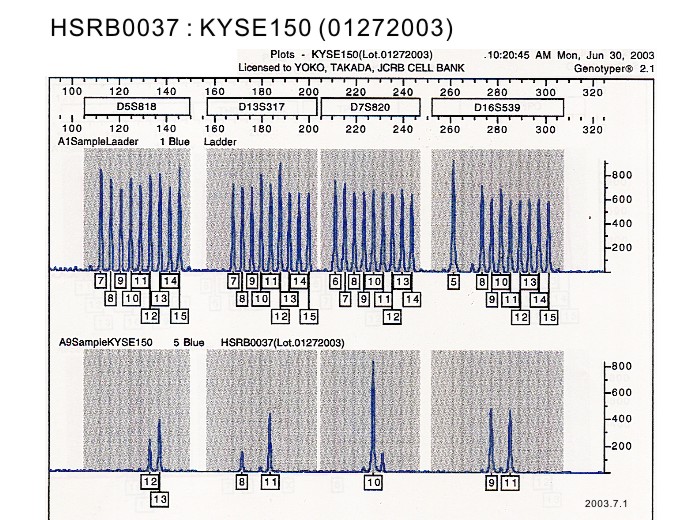
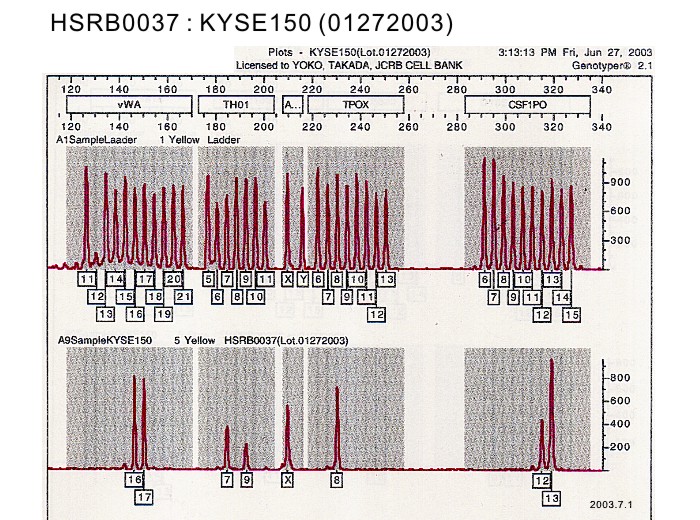
| Surface Antigen | DNA Profile (STR) | D5S818:12,13 D13S317:8,11 D7S820:10,11 D16S539:9,11 VWA:16,17 TH01:7,9 AM:X TPOX:8 CSF1PO:12,13 |
|
| Adhesion | Yes | Exoteric Gene | |
| Medium for Freezing | 10% DMSO, 20%& FBS - Ham's F12 | CO2 Conc. | 5% |
| Viability immediately after thawing (%) | Additional information |
| Cell No. | JCRB1095 | Cell Name | |
|---|---|---|---|
| LOT No. | 08272007 | Lot Specification | distribution |
| Medium | Ham's F12 medium with 2% fetal calf serum lotGIBCI484375 | Temperature | 37 |
| Cell Density at Seeding | 3-4 x10^4cells/ml | Methods for Passages | 0.25% trypsin and 0.02% EDTA in PBS |
| Doubling Time | 85hr | Cell Number in Vial (cells/1ml) | 2.8 x 10^6 cells |
| Viability at cell freezing (%) | 96 | Antibiotics Used | free |
| Passage Number | P6* | PDL | |
| Sterility: MYCOPLASMA | - | Sterility: BACTERIA | - |
| Sterility: FUNGI | - | Isozyme Analysis | Confirmed as human by NP, G6PD (type B) AST |
| Chromosome Mode | Chromosome Information | ||
| Surface Antigen | DNA Profile (STR) | ||
| Adhesion | Yes | Exoteric Gene | |
| Medium for Freezing | 10% DMSO, 20% FCS-culture medium | CO2 Conc. | 5% |
| Viability immediately after thawing (%) | Additional information |
| Cell No. | JCRB1095 | Cell Name | KYSE150 |
|---|---|---|---|
| LOT No. | 12102015 | Lot Specification | distribution |
| Medium | Ham's F12 medium(SIGMA,N6658) with 5% heat inactivated fetal bovine serum(SIGMA12J396) | Temperature | 37 C |
| Cell Density at Seeding | 0.3-1.0x10^5 cells/sq.cm | Methods for Passages | Cells were harvested after treatment with 0.25%trypsin(GIBCO) and 0.02% EDTA(1/3 split) once a week. |
| Doubling Time | NT | Cell Number in Vial (cells/1ml) | 2.6x10^6 |
| Viability at cell freezing (%) | 96.3 | Antibiotics Used | free |
| Passage Number | p10* | PDL | NT |
| Sterility: MYCOPLASMA | - | Sterility: BACTERIA | - |
| Sterility: FUNGI | - | Isozyme Analysis | NT |
| Chromosome Mode | NT | Chromosome Information | NT |
| Surface Antigen | NT | DNA Profile (STR) | D5S818:12,13 D13S317:8,11 D7S820:10,11 D16S539:9,11 VWA:16,17 TH01:7,9 AM:X TPOX:8 CSF1PO:12,13 |
| Adhesion | Yes | Exoteric Gene | NT |
| Medium for Freezing | BAMBANKER(LYMPHOTEC Inc.,CS-02-001,NIPPON Genetics Co., LTD) | CO2 Conc. | 5 % |
| Viability immediately after thawing (%) | Additional information |