JCRB1082 KYSE170
Cell information
Important Notice(s)Announcement of unavailability of KYSE series cell lines to oversea users outside of Japan
Cell type:general cells (View Pricing Information)
| JCRB No. | JCRB1082 | Cell Name | KYSE170 |
|---|---|---|---|
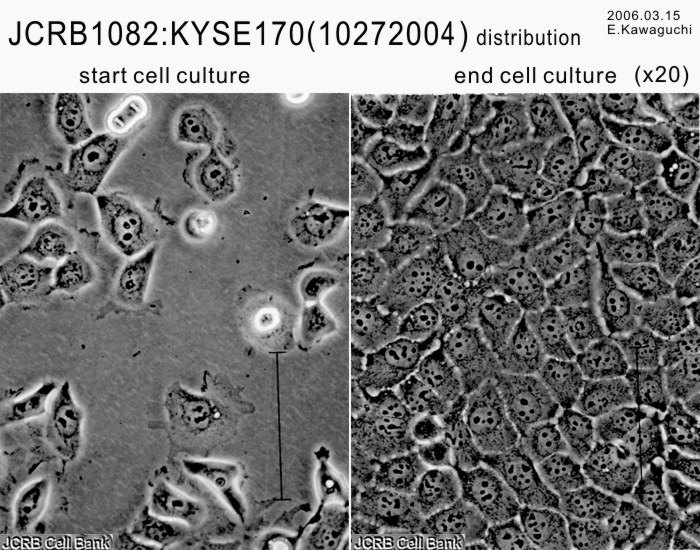
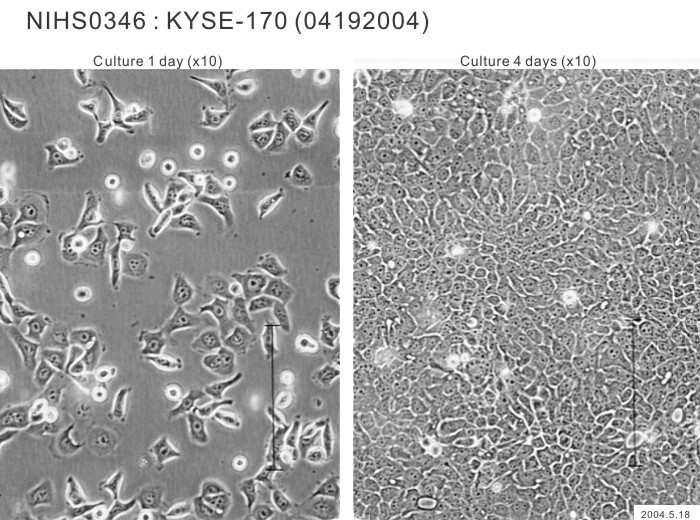
| Profile | Human squamous cell carcinoma cell line established from esophageal cancer. | Other Name | |
| Animal | human | Strain | |
| Genus | Homo | Species | sapiens |
| Sex | F | Age | 53 |
| Identity | available | Tissue for Primary Cancer | esophagus |
| Case history | esophagus cancer, T3N0M0 Stage 2a, squamous cell carcinoma (SCC) | Metastasis | No |
| Tissue Metastasized | Genetics | C-Myc and MDM2 amplification observed. p53 mutation, FHIT. | |
| Life Span | infinite | Crisis PDL | |
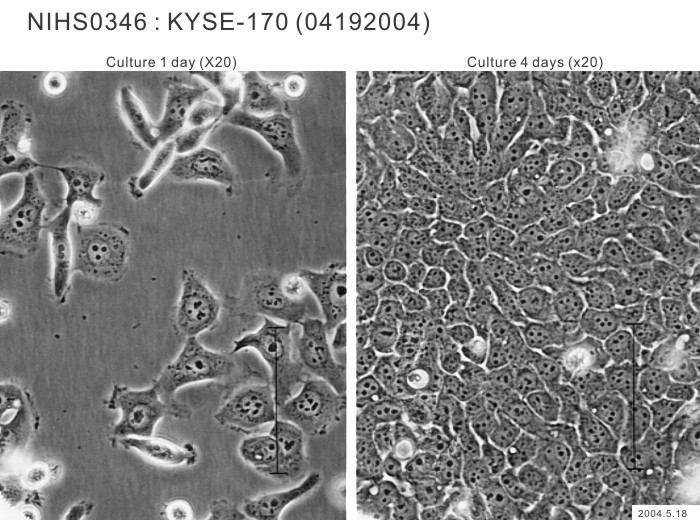
| Morphology | epithelial-like | Character | Xenotransplantation positive. |
| Classify | tumor | Established by | Shimada,Y. |
| Registered by | Shimada,Y. | Regulation for Distribution | |
| Comment | Year | 2003 | |
| Medium | Ham's F12 medium + RPMI1640 medium (1 to 1 mix) with 2% fetal calf serum. | Methods for Passages | Cells harvested after treatment with trypsin and EDTA. |
| Cell Number on Passage | split ratio = 1/5 | Race | Japanese |
| CO2 Conc. | 5 % | Tissue Sampling | esophagus |
| Tissue Type | mederately differenciated SCC |
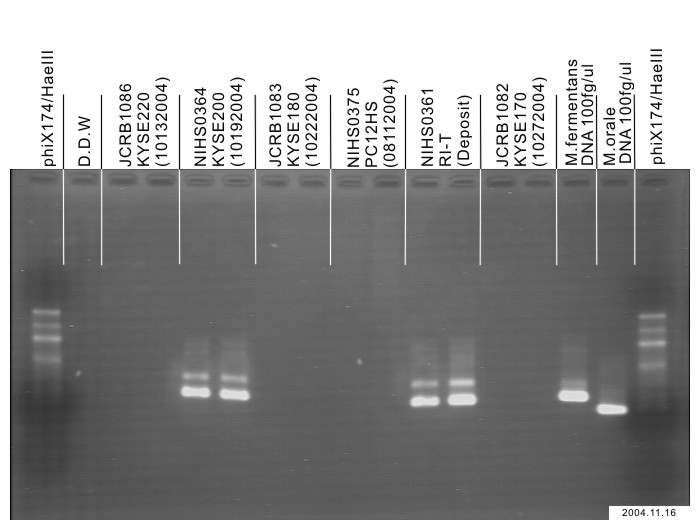
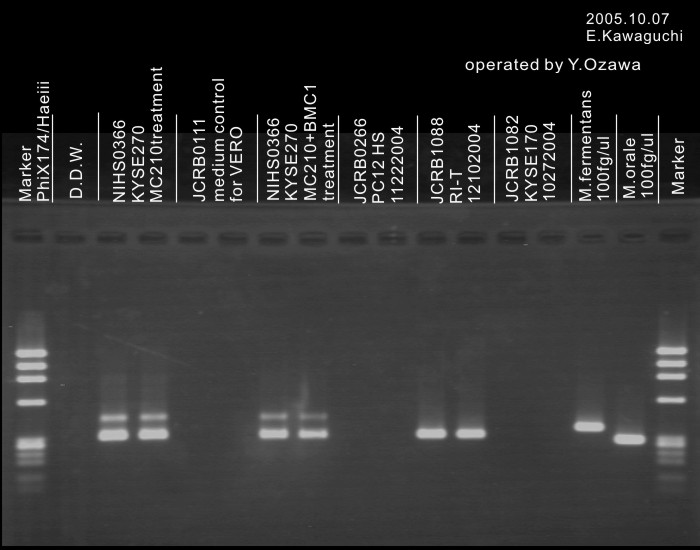
| Detection of virus genome fragment by Real-time PCR | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Detected DNA Virus | tested | Detected RNA Virus | tested | ||||||
| CMV | - | parvoB19 | - | HCV | - | HTLV-1 | - | ||
| EBV | - | HBV | - | HIV-1 | - | HTLV-2 | - | ||
| HHV6 | - | HTLV-1 | - | HIV-2 | - | HAV | - | ||
| HHV7 | - | HTLV-2 | - |
-/negative. +/positive. nt/not tested. (positive (+) does not immediately mean the production of infectious viral particles.) |
|||||
| BKV | - | HIV-1 | - | ||||||
| JCV | - | HIV-2 | - | ||||||
| ADV | - | HPV18 | - | ||||||
| Notes | |||||||||
| Reference | |
|---|---|
| Pubmed id:11092977 | Nonrandom chromosomal imbalances in esophageal squamous cell carcinoma cell lines: possible involvement of the ATF3 and CENPF genes in the 1q32 amplicon. Pimkhaokham A,Shimada Y,Fukuda Y,Kurihara N,Imoto I,Yang ZQ,Imamura M,Nakamura Y,Amagasa T,Inazawa J Jpn J Cancer Res. 2000 Nov;91(11):1126-33 |
| Pubmed id:10446988 | Lymph node metastasis is associated with allelic loss on chromosome 13q12-13 in esophageal squamous cell carcinoma. Harada H,Tanaka H,Shimada Y,Shinoda M,Imamura M,Ishizaki K Cancer Res. 1999 Aug 1;59(15):3724-9 |
| Pubmed id:9699676 | Methylation of the 5' CpG island of the FHIT gene is closely associated with transcriptional inactivation in esophageal squamous cell carcinomas. Tanaka H,Shimada Y,Harada H,Shinoda M,Hatooka S,Imamura M,Ishizaki K Cancer Res. 1998 Aug 1;58(15):3429-34 |
| Pubmed id:9033652 | Multiple types of aberrations in the p16 (INK4a) and the p15(INK4b) genes in 30 esophageal squamous-cell-carcinoma cell lines. Tanaka H,Shimada Y,Imamura M,Shibagaki I,Ishizaki K Int J Cancer. 1997 Feb 7;70(4):437-42 |
| Pubmed id:8575860 | Characterization of p53 gene mutations in esophageal squamous cell carcinoma cell lines: increased frequency and different spectrum of mutations from primary tumors. Tanaka H,Shibagaki I,Shimada Y,Wagata T,Imamura M,Ishizaki K Int J Cancer. 1996 Jan 26;65(3):372-6 |
| Pubmed id:9816044 | p53 mutation, murine double minute 2 amplification, and human papillomavirus infection are frequently involved but not associated with each other in esophageal squamous cell carcinoma. Shibagaki I,Tanaka H,Shimada Y,Wagata T,Ikenaga M,Imamura M,Ishizaki K Clin Cancer Res. 1995 Jul;1(7):769-73 |
| Pubmed id:7913084 | Analysis of gene amplification and overexpression in human esophageal-carcinoma cell lines. Kanda Y,Nishiyama Y,Shimada Y,Imamura M,Nomura H,Hiai H,Fukumoto M Int J Cancer. 1994 Jul 15;58(2):291-7 |
| Pubmed id:8518899 | Prognostic significance of cell culture in carcinoma of the oesophagus. Shimada Y,Imamura M Br J Surg. 1993 May;80(5):605-7 |
| Pubmed id:1728357 | Characterization of 21 newly established esophageal cancer cell lines. Shimada Y,Imamura M,Wagata T,Yamaguchi N,Tobe T Cancer. 1992 Jan 15;69(2):277-84 |
| Images |
|---|
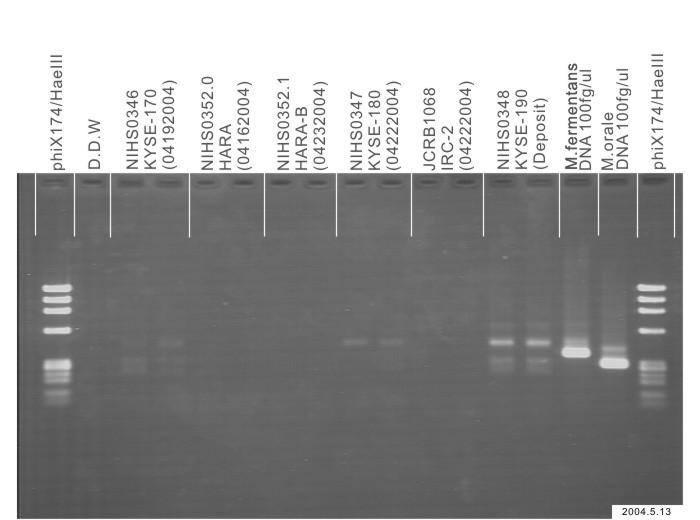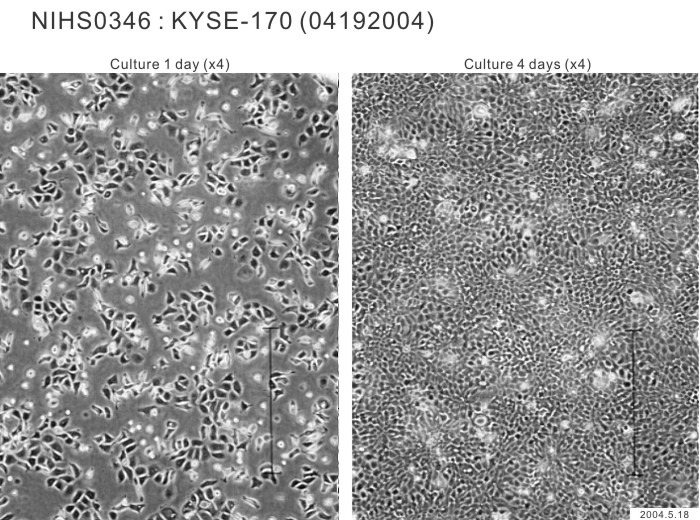                   |
LOT Information
Viability/Growth rate/Cell number are represented as actual values measured at lot presentation in JCRB, but are not guaranteed values. Additionally, the doubling time is a rough value measured during passages.| Cell No. | JCRB1082 | Cell Name | KYSE170 |
|---|---|---|---|
| LOT No. | 10272004 | Lot Specification | distribution |
| Medium | RPMI1640 and Ham's F12 medium(1/1) with 2% heat inactivated fetal bovine serum | Temperature | 37 C |
| Cell Density at Seeding | 4.5x10^4 | Methods for Passages | 0.25% trypsin and 0.02% EDTA |
| Doubling Time | NT | Cell Number in Vial (cells/1ml) | 2.7x10^6 |
| Viability at cell freezing (%) | 93.2 | Antibiotics Used | Free |
| Passage Number | P691 | PDL | NT |
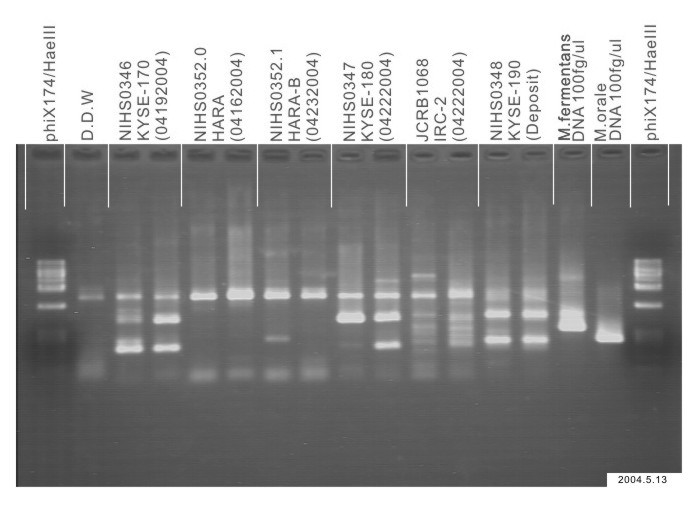
| Sterility: MYCOPLASMA | - | Sterility: BACTERIA | NT |
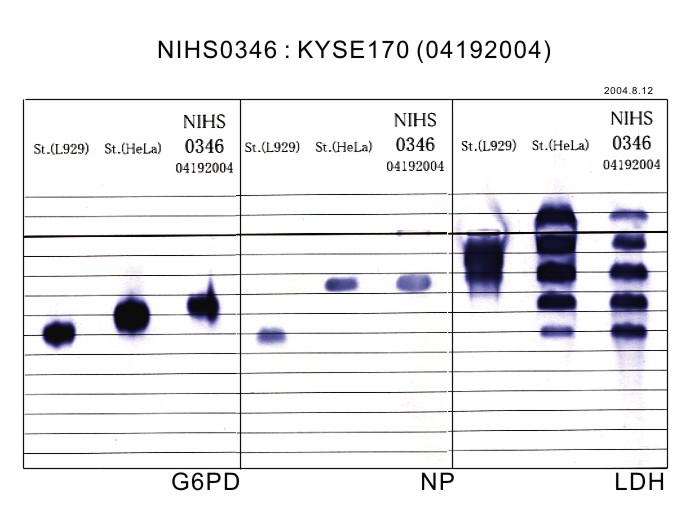
| Sterility: FUNGI | NT | Isozyme Analysis | NT |
| Chromosome Mode | NT | Chromosome Information | NT |
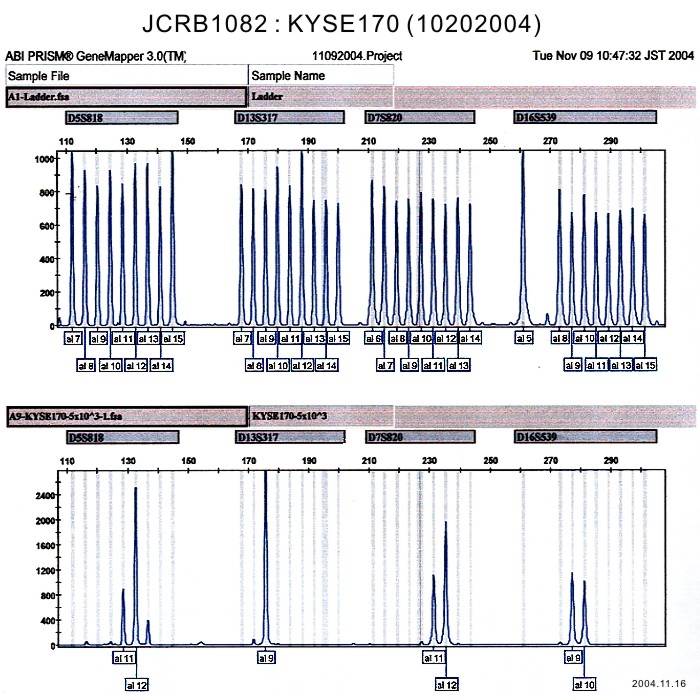
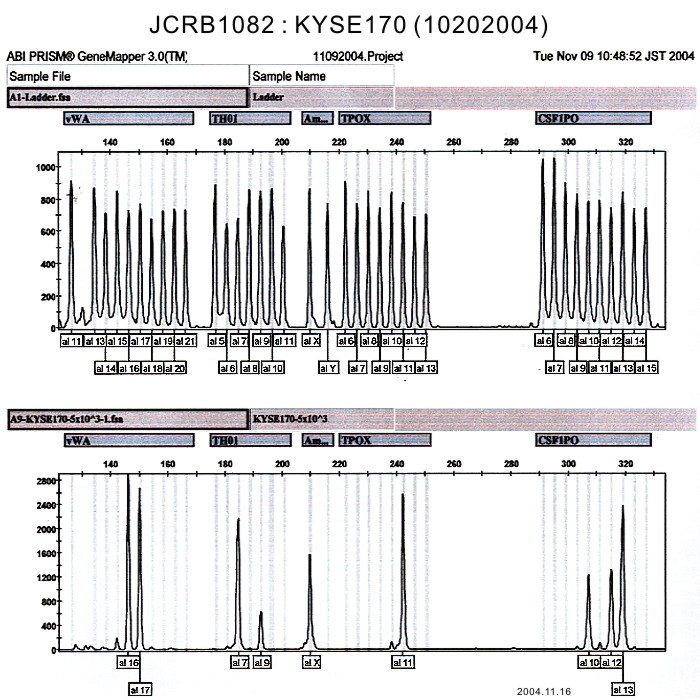
| Surface Antigen | NT | DNA Profile (STR) | |
| Adhesion | Yes | Exoteric Gene | NT |
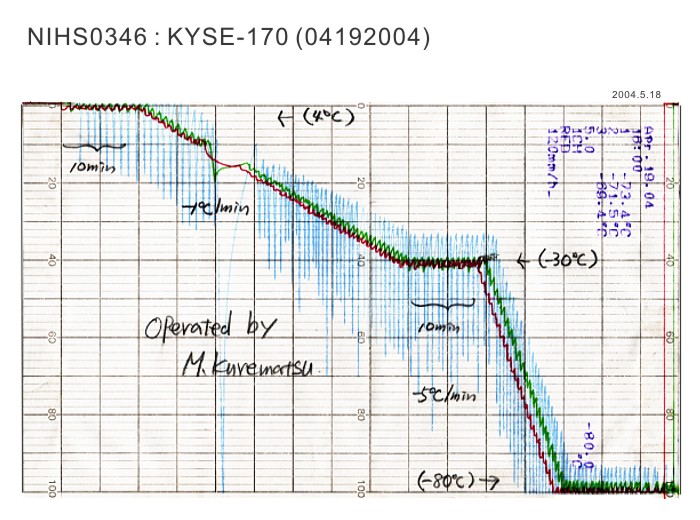
| Medium for Freezing | Culture medium with final 10% fetal bovine serum and 5% DMSO | CO2 Conc. | 5 % |
| Viability immediately after thawing (%) | Additional information |
| Images |
|---|
      |
| Cell No. | JCRB1082 | Cell Name | KYSE170 |
|---|---|---|---|
| LOT No. | 10052015 | Lot Specification | distribution |
| Medium | 1:1 mixture of Ham's F12 medium and RPMI1640 medium with 2% heat-inactivated fetal bovine serum (FBS; GIBCO Cat. # 10091). | Temperature | 37 C |
| Cell Density at Seeding | 5.8 - 9.8 x 10^5 cells/mL | Methods for Passages | Cells were harvested after treatment with 0.25% trypsin and 0.02% EDTA. |
| Doubling Time | 19 - 29 hrs | Cell Number in Vial (cells/1ml) | 2.4 x 10^6 |
| Viability at cell freezing (%) | 96.8 | Antibiotics Used | free |
| Passage Number | P690 | PDL | |
| Sterility: MYCOPLASMA | - | Sterility: BACTERIA | - |
| Sterility: FUNGI | - | Isozyme Analysis | |
| Chromosome Mode | Chromosome Information | ||
| Surface Antigen | DNA Profile (STR) | D5S818:11,12,13 D13S317:9 D7S820:11,12 D16S539:9,10 VWA:16,17 TH01:7,9 AM:X TPOX:11 CSF1PO:10,12,13 |
|
| Adhesion | Yes | Exoteric Gene | |
| Medium for Freezing | 10% DMSO, 20% FBS - culture medium | CO2 Conc. | 5% |
| Viability immediately after thawing (%) | 96.8 | Additional information |